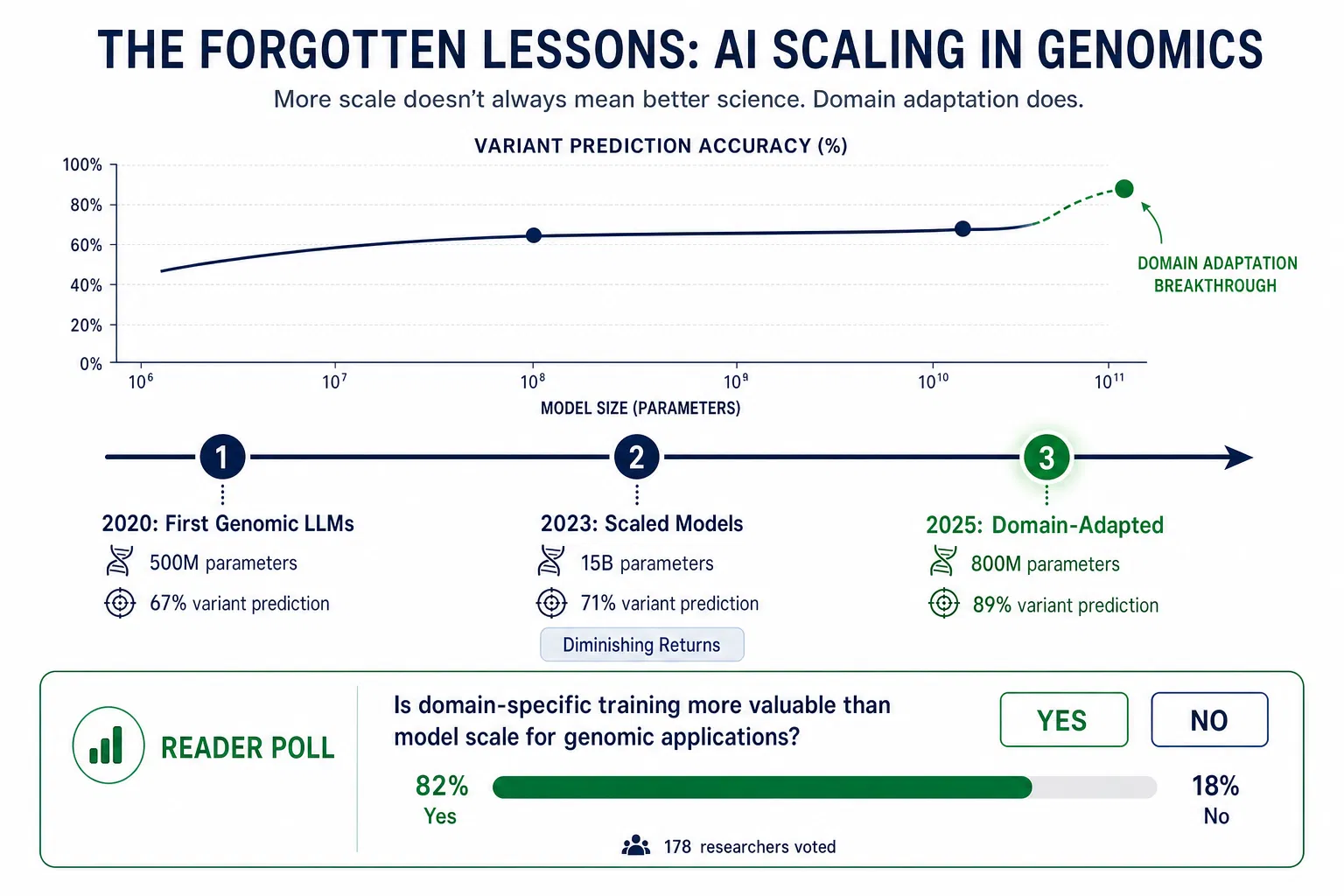
The dominant paradigm in artificial intelligence for the past decade has been remarkably simple in its logic: more data, more parameters, more compute, better results. Sara Hooker calls this the era of brute force, and her work demonstrates convincingly that this approach is approaching its limits — not merely in terms of cost and energy consumption, but in terms of fundamental capability. Nowhere are these limits more apparent than in genomic medicine, where the sheer complexity of biological systems, the heterogeneity of patient populations, and the rapid pace of new discoveries demand an entirely different kind of intelligence. Not bigger models, but smarter ones. Not static systems frozen at the moment of training, but adaptive systems that evolve alongside the science they serve.
The human genome contains approximately three billion base pairs, but this number vastly understates the complexity that genomic medicine must navigate. Epigenetic modifications, alternative splicing, post-translational modifications, gene-gene interactions, gene-environment interactions, and the dynamic regulation of gene expression across tissues and developmental stages create a combinatorial space that no amount of brute-force scaling can adequately model. Traditional large language models trained on genomic sequences can identify statistical patterns in DNA, but they lack the adaptive capacity to integrate new biological knowledge as it emerges — a critical limitation in a field where our understanding of genome function is revised almost daily.
Hooker's research on efficient, adaptive AI systems offers a fundamentally different approach. Rather than building monolithic models that attempt to encode all genomic knowledge at the moment of training, adaptive systems can continuously incorporate new findings — newly characterized regulatory elements, recently discovered disease-associated variants, emerging understanding of non-coding RNA function — without the prohibitive cost of retraining from scratch. This capability is not merely a computational convenience; it is a prerequisite for any AI system that aspires to support clinical genomic interpretation in real time.
The pharmacogenomics application illustrates this principle with particular clarity. Individual patients respond differently to medications based on their genetic variants in drug-metabolizing enzymes, drug transporters, and drug targets. The catalog of clinically actionable pharmacogenomic variants is expanding rapidly, with new associations published weekly. A static AI system, no matter how large, cannot keep pace with this evolving knowledge base. An adaptive system, by contrast, can integrate each new pharmacogenomic discovery as it emerges, maintaining clinical relevance without requiring complete retraining — a process that for large models can cost millions of dollars and weeks of computation.
Hooker's concept of the hardware lottery also resonates deeply in genomic medicine. The computational architectures that dominate AI research were designed for tasks with fundamentally different properties than genomic analysis. Image classification and natural language processing operate on data with relatively uniform structure and well-defined boundaries. Genomic data, by contrast, exhibits long-range dependencies spanning millions of base pairs, hierarchical organization across multiple scales, and context-dependent interpretation that varies by tissue type, developmental stage, and environmental condition. Building AI systems that can navigate this complexity requires not just algorithmic innovation but architectural innovation — new computational substrates designed with biological data in mind.
The equity implications of adaptive AI in genomic medicine deserve particular attention. Current genomic databases are overwhelmingly derived from populations of European descent, creating systematic biases in variant interpretation and polygenic risk scores. Adaptive AI systems that can learn efficiently from limited data offer a path toward more equitable genomic medicine — systems that can provide accurate interpretation for underrepresented populations without requiring the massive datasets that have historically been available only for well-studied groups. This aligns with Hooker's broader vision of AI that serves diverse populations rather than optimizing for the average case.
The convergence of adaptive AI and genomic medicine represents one of the most promising frontiers in contemporary science. As our understanding of genome function deepens and the cost of genomic sequencing continues to fall, the bottleneck increasingly lies not in data generation but in intelligent interpretation. Hooker's work provides both the theoretical framework and the practical methodology for building AI systems equal to this challenge — systems that are efficient enough to deploy at scale, adaptive enough to evolve with new knowledge, and equitable enough to serve all patients regardless of their genetic ancestry. The future of genomic medicine is not bigger models; it is smarter, more adaptive ones.