The Silent Crisis in African Grain
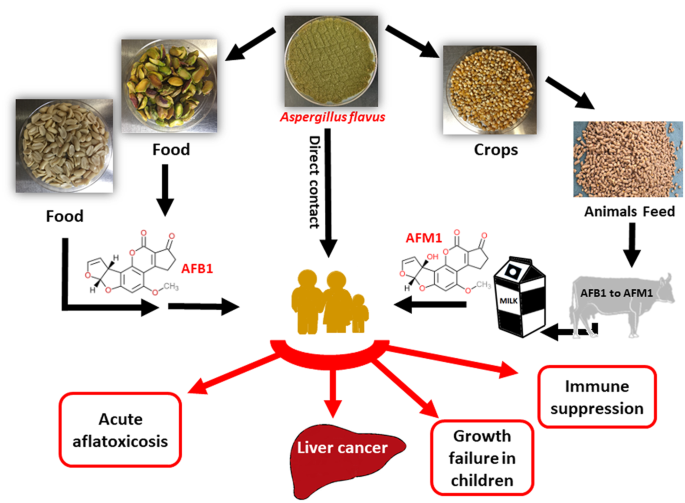
Every year, across the maize fields of Kenya, the groundnut farms of Nigeria, and the sorghum stores of Ethiopia, a microscopic fungus conducts a slow, invisible assault on the food supply of hundreds of millions of people. Aspergillus flavus — a ubiquitous soil-dwelling mould — colonises cereal grains, legumes, and oilseeds, producing aflatoxins: a family of polyketide-derived secondary metabolites that are among the most potent naturally occurring carcinogens known to science.
The consequences are devastating and largely invisible to global health statistics. Aflatoxin B1 (AFB1), the most toxic and prevalent of the aflatoxin family, is classified as a Group 1 human carcinogen by the International Agency for Research on Cancer. Chronic dietary exposure to AFB1 is the primary environmental risk factor for hepatocellular carcinoma (HCC) — liver cancer — in Sub-Saharan Africa and Southeast Asia. In combination with hepatitis B virus infection, AFB1 exposure increases HCC risk by a factor of 30 or more, creating a synergistic carcinogenic burden that disproportionately affects the world's most food-insecure populations.
Beyond cancer, aflatoxin exposure causes acute aflatoxicosis — a potentially fatal condition characterised by liver failure, haemorrhage, and immune suppression — and chronic subclinical effects including stunted growth in children, immune dysfunction, and reduced vaccine efficacy. The African Union estimates that mycotoxin contamination costs the continent approximately US$4 billion annually in lost agricultural trade.
The Molecular Biology of Aflatoxin Production
Understanding why A. flavus produces aflatoxins — and under what conditions — is the foundation for any effective intervention strategy. Aflatoxin biosynthesis is governed by a 29-gene cluster located on chromosome 3 of A. flavus, encoding a complex polyketide pathway that converts acetyl-CoA and malonyl-CoA into the mature aflatoxin molecule through approximately 23 enzymatic steps.
The pathway is regulated by a master transcription factor, AflR, which activates the entire biosynthetic cluster in response to specific environmental signals: nitrogen limitation, carbon source availability, temperature stress (optimal production occurs between 25–35°C), water activity (optimal between 0.95–0.99 aw), and oxidative stress. The pathway is also regulated by the global regulator LaeA, which controls secondary metabolism across the Aspergillus genus through histone methylation, and by the velvet complex (VeA, VelB, VosA), which integrates light, temperature, and developmental signals.
This regulatory complexity has important practical implications. Aflatoxin contamination is not a fixed property of a grain lot — it is a dynamic outcome of the interaction between fungal genetics, host plant physiology, and environmental conditions. This dynamic, context-dependent nature is precisely what makes aflatoxin contamination so difficult to manage with conventional surveillance approaches — and precisely what makes it amenable to AI-based predictive modelling.
Molecular Epidemiology: Mapping the Diversity of Aflatoxigenic Strains
Not all A. flavus isolates are equally dangerous. The species is divided into two morphological groups — S-strain (small sclerotia, uniformly high aflatoxin production) and L-strain (large sclerotia, variable aflatoxin production) — and within these groups, there is enormous genetic diversity in aflatoxin production capacity. Some A. flavus isolates are atoxigenic — they carry the biosynthetic gene cluster but produce little or no aflatoxin due to mutations in key pathway genes.
Atoxigenic strains of A. flavus have been developed as biological control agents. When applied to fields at high density, they competitively exclude toxigenic strains and dramatically reduce aflatoxin contamination. The Aflasafe product family, developed by the International Institute of Tropical Agriculture (IITA), has demonstrated reductions in aflatoxin contamination of 80–99% in field trials across multiple African countries.
However, the effectiveness of biocontrol depends critically on the genetic composition of the local A. flavus population. Whole-genome sequencing of A. flavus isolates from across Sub-Saharan Africa, combined with machine learning-based population structure analysis, can identify the dominant toxigenic lineages in each agroecological zone, predict their competitive dynamics with candidate biocontrol strains, and guide the selection and deployment of regionally optimised biocontrol products.
AI-Powered Predictive Surveillance: From Field to Fork
The conventional approach to aflatoxin surveillance relies on physical sampling of grain lots at key points in the supply chain, followed by laboratory analysis using immunoassay or chromatographic methods. This approach is expensive, slow, and spatially sparse: it can detect contamination after it has occurred but cannot predict where and when contamination will emerge.
AI-based predictive surveillance offers a fundamentally different approach. By integrating multiple data streams — satellite remote sensing, weather station data, soil moisture sensors, crop health indices, historical contamination records, and molecular epidemiological data — machine learning models can generate spatially explicit, temporally dynamic risk maps that predict aflatoxin contamination risk at the field level, weeks before harvest.
| Data Type | Source | Relevance to Aflatoxin Risk |
|---|---|---|
| Land surface temperature | MODIS, Landsat | Drives A. flavus growth and AflR activation |
| NDVI (crop health index) | Sentinel-2 | Proxy for crop stress, which increases host susceptibility |
| Soil moisture | SMAP satellite, ground sensors | Determines water activity in grain, critical for fungal growth |
| Rainfall anomalies | CHIRPS, weather stations | Drought stress increases susceptibility; post-harvest rain increases storage risk |
| Historical contamination records | National food safety agencies | Provides ground truth for model training and validation |
| A. flavus population density | Soil sampling, spore trapping | Direct measure of inoculum pressure |
Random forest and gradient boosting models have been used to predict contamination risk at the district level using satellite-derived environmental variables, achieving AUC values of 0.85–0.92 in cross-validation studies. Convolutional neural networks applied to multispectral satellite imagery have demonstrated the ability to identify pre-harvest crop stress patterns associated with elevated contamination risk. LSTM networks have been used to model the temporal dynamics of contamination risk across the growing season.
The One Health Dimension: Connecting Grain, Animal, and Human Health
Aflatoxin contamination does not respect the boundaries between agricultural, veterinary, and human health systems. The same contaminated grain that poses a cancer risk to human consumers also enters the livestock feed supply, where aflatoxin M1 (a hydroxylated metabolite of AFB1) is secreted into milk and can reach human consumers through dairy products.
This interconnection demands a One Health approach to aflatoxin surveillance and management — one that integrates data from human health surveillance (HCC incidence, acute aflatoxicosis cases), veterinary monitoring (aflatoxin M1 in milk, poultry performance indicators), and agricultural monitoring (grain contamination, crop stress indices) into a unified analytical framework. AI-based data integration platforms are uniquely suited to this task, connecting databases that currently operate in institutional silos.
Policy Implications: Building an African Aflatoxin Governance Framework
The current regulatory landscape for aflatoxin in Sub-Saharan Africa is fragmented and inconsistent. Maximum residue limits (MRLs) for aflatoxin in food vary widely across countries — from 4 ppb (total aflatoxins) in Kenya and South Africa, aligned with European Union standards, to 20 ppb or higher in several other African countries. This regulatory heterogeneity creates perverse incentives: grain that fails to meet the stringent standards of export markets is diverted to domestic markets, where it is consumed by the most vulnerable populations.
A coherent African aflatoxin governance framework — one that harmonises MRLs across the continent, establishes shared surveillance infrastructure, and integrates aflatoxin risk into national food security early warning systems — is an urgent policy priority. The African Union's CAADP and the AfCFTA provide institutional platforms for this harmonisation.
Conclusion: A Research Agenda for the Next Decade
The convergence of molecular biology, AI, and policy science offers a genuine opportunity to transform aflatoxin management in Sub-Saharan Africa. The research agenda for the next decade should prioritise four interconnected objectives: comprehensive molecular epidemiological mapping of A. flavus populations across African agroecological zones; development and validation of AI-based predictive surveillance systems; construction of One Health data integration platforms; and policy research on governance frameworks for aflatoxin management at the scale of the African Continental Free Trade Area.
This is not a distant aspiration. The data, the methods, and the institutional platforms needed to pursue this agenda exist today. What is required is the will to invest in knowledge architecture — the ontologies, data pipelines, and governance frameworks that transform fragmented scientific data into actionable intelligence — and the recognition that food security in Africa is, at its core, a biosecurity challenge that demands the full integration of molecular science, computational intelligence, and policy leadership.