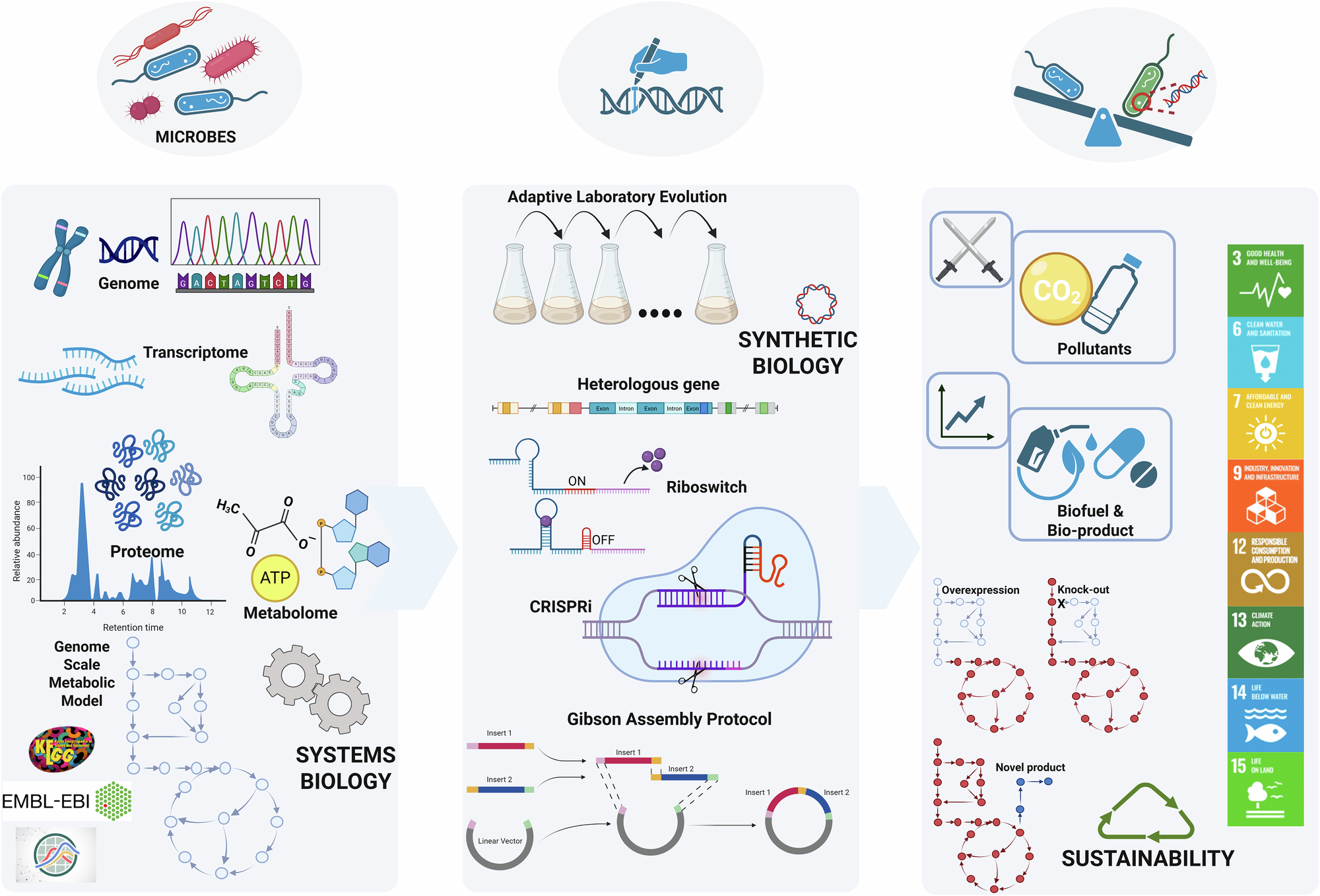
Microorganisms are, at their core, extraordinarily sophisticated chemical factories. Through billions of years of evolution, bacteria, yeasts, and fungi have developed metabolic networks of staggering complexity — networks capable of converting simple carbon sources into an enormous diversity of organic compounds. The challenge for metabolic engineers has always been to redirect these networks toward the production of compounds that are useful to humanity: pharmaceuticals, biofuels, biodegradable plastics, flavour compounds, and industrial enzymes. The challenge is formidable because metabolic networks are not linear pipelines but densely interconnected webs of reactions, each influencing the others in ways that are difficult to predict from first principles.
Artificial intelligence is fundamentally changing the way metabolic engineers approach this challenge. By combining genome-scale metabolic models, machine learning algorithms, and high-throughput experimental data, researchers are now able to design, predict, and optimise microbial production strains with a speed and precision that was unimaginable a decade ago. Nowhere is this more consequential than in the production of polyhydroxyalkanoates (PHAs) — a family of biodegradable polyesters that represent one of the most promising alternatives to petroleum-derived plastics.
The PHA Problem: Promise and Bottleneck
Polyhydroxyalkanoates are naturally produced by a wide range of bacteria as intracellular carbon and energy storage granules under conditions of nutrient limitation. They are fully biodegradable, biocompatible, and can be produced from renewable feedstocks — making them an attractive alternative to conventional plastics in packaging, medical devices, and agricultural films. The global PHA market is projected to reach USD 135 million by 2027, driven by tightening regulations on single-use plastics and growing consumer demand for sustainable materials.
The primary barrier to widespread PHA adoption is cost. Conventional petroleum-derived plastics are produced at a fraction of the cost of microbially synthesised PHAs, largely because the metabolic yields of current production strains are insufficient to achieve economic competitiveness. Improving those yields requires optimising a complex web of metabolic decisions: which carbon sources to use, which competing pathways to downregulate, which regulatory genes to modify, and how to balance the competing demands of cell growth and polymer accumulation. This is precisely the kind of multi-dimensional optimisation problem at which AI excels.
Genome-Scale Metabolic Models and Flux Balance Analysis
The foundation of AI-driven metabolic engineering is the genome-scale metabolic model (GEM) — a mathematical representation of all the metabolic reactions encoded in an organism's genome, constrained by stoichiometry and thermodynamics. GEMs allow researchers to simulate the metabolic behaviour of an organism under different conditions and genetic configurations, predicting which genetic modifications are most likely to increase the flux of carbon toward a target product.
The most widely used computational method for analysing GEMs is flux balance analysis (FBA), which uses linear programming to find the distribution of metabolic fluxes that maximises a defined objective function — typically biomass production or product yield — subject to the stoichiometric and capacity constraints of the network. FBA has been used to identify gene knockout targets for PHA overproduction in organisms including Cupriavidus necator (formerly Ralstonia eutropha), Pseudomonas putida, and engineered strains of Escherichia coli.
| Organism | Native PHA Type | AI/ML Application | Reported Yield Improvement |
|---|---|---|---|
| Cupriavidus necator H16 | PHB (poly-3-hydroxybutyrate) | FBA + reinforcement learning for gene knockout | Up to 40% increase in PHB content |
| Pseudomonas putida KT2440 | mcl-PHA | Deep learning for promoter optimisation | 2.3-fold increase in titre |
| E. coli (engineered) | PHB, PHBV | ML-guided combinatorial pathway design | 85% of theoretical maximum yield |
| Haloferax mediterranei | PHBV | Genome-scale modelling + flux sampling | Identification of 12 novel knockout targets |
| Synechocystis sp. PCC 6803 | PHB (photoautotrophic) | Neural network for light/nutrient optimisation | 3.1-fold increase under optimised conditions |
Machine Learning for Enzyme Engineering and Pathway Design
Beyond flux optimisation, machine learning is being applied to the engineering of the individual enzymes that constitute PHA biosynthetic pathways. The key enzymes — PhaA (β-ketothiolase), PhaB (acetoacetyl-CoA reductase), and PhaC (PHA synthase) — have been extensively studied, but their activity, substrate specificity, and stability can be further improved through directed evolution guided by machine learning.
Protein language models such as ESM-2 and ProtTrans, trained on hundreds of millions of protein sequences, can predict the functional consequences of amino acid substitutions with remarkable accuracy — enabling researchers to design enzyme variants with improved catalytic efficiency without the need for exhaustive experimental screening. In a landmark 2023 study, researchers used a variational autoencoder trained on PhaC synthase sequences to generate novel enzyme variants with up to 4.7-fold higher activity than the wild-type enzyme, identifying mutations in regions of the protein that had not previously been targeted by rational design.
Reinforcement learning is also being applied to the higher-level problem of pathway design — selecting which genes to express, at what levels, and in which cellular compartment, to maximise product yield while minimising metabolic burden. These approaches treat the metabolic engineering problem as a sequential decision-making task, with the reinforcement learning agent learning from the outcomes of iterative experimental rounds to converge on optimal genetic configurations.
The Design-Build-Test-Learn Cycle Accelerated
The traditional metabolic engineering workflow — hypothesise a genetic modification, construct the strain, measure the phenotype, interpret the results — is inherently slow and expensive. Each cycle can take weeks to months, and the number of possible genetic configurations is astronomically large. AI is accelerating this cycle at every stage.
At the design stage, generative models can propose novel genetic configurations that human engineers would not have considered. At the build stage, automated DNA synthesis and assembly platforms can construct dozens of strain variants in parallel. At the test stage, high-throughput metabolomics and proteomics platforms can characterise the metabolic phenotype of each variant rapidly. At the learn stage, machine learning models trained on the accumulated experimental data can identify the genetic features most predictive of high PHA yield, guiding the next design cycle with increasing precision.
This closed-loop approach — sometimes called the Design-Build-Test-Learn (DBTL) cycle — is being implemented at scale by companies including Zymergen (now part of Ginkgo Bioworks), Ginkgo Bioworks itself, and Amyris. The results are striking: what once required years of manual strain engineering can now be accomplished in months, with AI-guided strain libraries exploring a vastly larger design space than any human team could navigate alone.
Beyond PHA: The Broader Metabolic Engineering Landscape
The AI-driven metabolic engineering approaches developed for PHA production are being applied across the full spectrum of bioproduction targets. In the pharmaceutical sector, AI-guided pathway engineering has been used to optimise the production of artemisinin (an antimalarial compound), taxol precursors, and opioid biosynthetic intermediates in engineered yeast. In the food and flavour industry, machine learning has guided the engineering of E. coli strains for the production of vanillin, lycopene, and resveratrol. In the biofuel sector, AI-optimised strains of Clostridium and Saccharomyces are achieving ethanol and butanol yields that approach the theoretical thermodynamic limits.
The convergence of AI, synthetic biology, and metabolic engineering is creating a new paradigm for the production of materials and chemicals — one in which the design space is explored computationally rather than empirically, and in which the pace of innovation is limited not by the speed of laboratory experimentation but by the quality of the models and the data that train them. For a world facing the twin challenges of climate change and plastic pollution, this convergence could not be more timely.