In the classical preclinical research workflow, generating a comprehensive pharmacological profile for a novel drug candidate requires months of animal experiments, specialist laboratory infrastructure, and significant financial investment. A single DART (dose, absorption, response, toxicity) study in a murine model — covering dose-response relationships, multi-organ toxicity, and biomarker kinetics — can cost tens of thousands of dollars and require the sacrifice of dozens of animals. For academic researchers and small biotechnology companies, this cost is often prohibitive.
Invitron has built an alternative: the DART Analysis framework, a computational pipeline that generates the same categories of preclinical data — dose-response curves, organ toxicity profiles, biomarker timelines, pharmacokinetic parameters, and translational scoring — entirely in silico, in minutes, and at a fraction of the cost. The framework is compliant with the FDA Modernization Act 2.0, aligned with ICH S7A/S7B cardiac safety guidelines, and built on validated digital equivalents of 16 research species.
The Four-Stage Pipeline
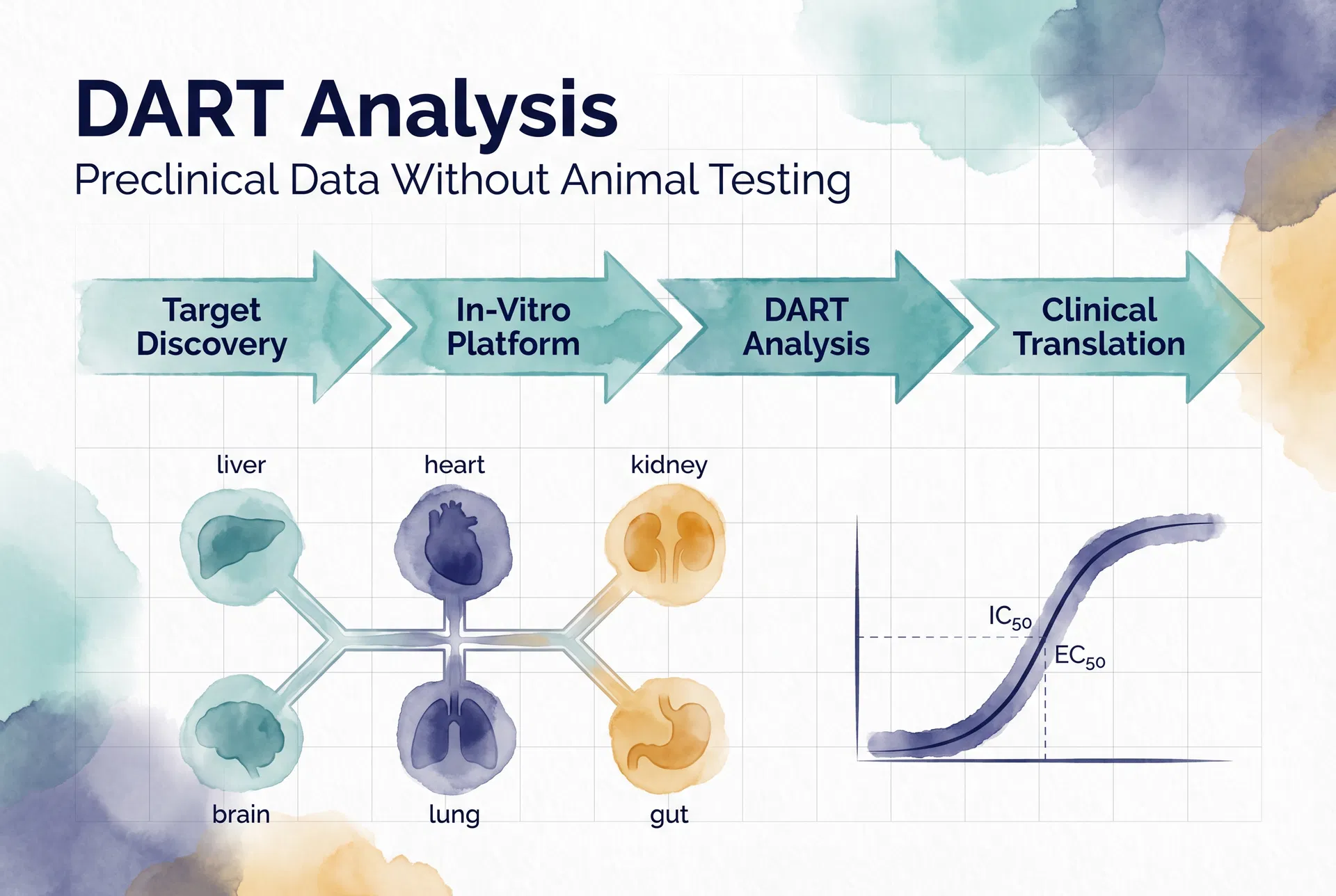
DART Analysis on Invitron proceeds through four sequential stages, each corresponding to a phase of the classical drug development workflow.
Stage 1 — Target Discovery establishes the biological context: the researcher selects a species (e.g., C57BL/6J mouse), a disease model (e.g., oncology — tumour growth and metastasis), and a drug compound (e.g., Imatinib, IC₅₀: 0.025 μM). The platform retrieves the relevant genetic background, disease-associated biomarker profiles, and baseline pharmacokinetic parameters for the selected combination.
Stage 2 — In-Vitro Platform simulates cellular-level drug interactions, generating predicted IC₅₀ and EC₅₀ values across the platform's organ-on-chip model. The six-organ system — liver (ALT, AST), heart (Troponin I, BNP), kidney (creatinine, BUN), brain (BDNF, NSE), lung (SP-D, KL-6), and gut (Zonulin, I-FABP) — provides a multi-organ toxicity profile that identifies off-target effects that single-organ studies routinely miss.
Stage 3 — DART Analysis is the computational core of the pipeline. It generates nine chart types: dose-response curves, 28-day biomarker timelines, organ toxicity heat maps, translational scoring matrices, pharmacokinetic profiles (IC₅₀/EC₅₀/LD₅₀), therapeutic index calculations, ML algorithm performance comparisons, feature importance rankings, and predictive confidence intervals. All nine charts are exportable in CSV, JSON, or formatted report output — ready for inclusion in a regulatory submission or peer-reviewed publication.
Stage 4 — Clinical Translation applies AI-powered human predictability scoring, estimating the likelihood that the observed in silico response will translate to human clinical outcomes. This translational scoring is calibrated against historical clinical trial data and is one of the most scientifically significant features of the platform — directly addressing the 90% attrition rate that plagues the transition from animal studies to human trials.
Machine Learning Enhancement
A key differentiator of Invitron's DART framework is its integration with the ML Periodic Table and Upload & Train modules. Researchers are not limited to the platform's pre-trained base models. By uploading their own experimental data — even a modest CSV file of dose-response observations from a cell-based assay — they can retrain the platform's predictive models using any of 40+ algorithms, producing a personalised DART analysis that reflects their specific experimental context.
The LoRA Studio module extends this further, enabling low-rank adaptation fine-tuning of the platform's biological prediction models with minimal data requirements. This is particularly valuable for researchers working with rare disease models or novel compound classes for which large training datasets do not yet exist.
Practical Implications for Researchers
For a PhD student or early-career researcher in a resource-limited setting, Invitron's DART framework changes what is possible. A researcher investigating a novel antimalarial compound can, within a single session on https://www.invitron.org, generate a complete preclinical pharmacological profile — species selection, strain engineering, disease modelling, dose-response analysis, multi-organ toxicity profiling, and clinical translation scoring — that would previously have required a fully equipped animal facility, an ethics committee approval, and six months of experimental work.
The platform's token-based access model, powered by the josephodongo.com Token Network, makes this accessible to researchers worldwide, including those in institutions that lack the infrastructure for conventional preclinical research. In a global research landscape where the gap between well-resourced and under-resourced institutions remains vast, tools like Invitron represent a meaningful step toward equity in scientific capability.
For researchers, educators, and drug developers ready to explore the next generation of preclinical research tools, Invitron is available at https://www.invitron.org.