The Fragmentation Problem in Bioinformatics
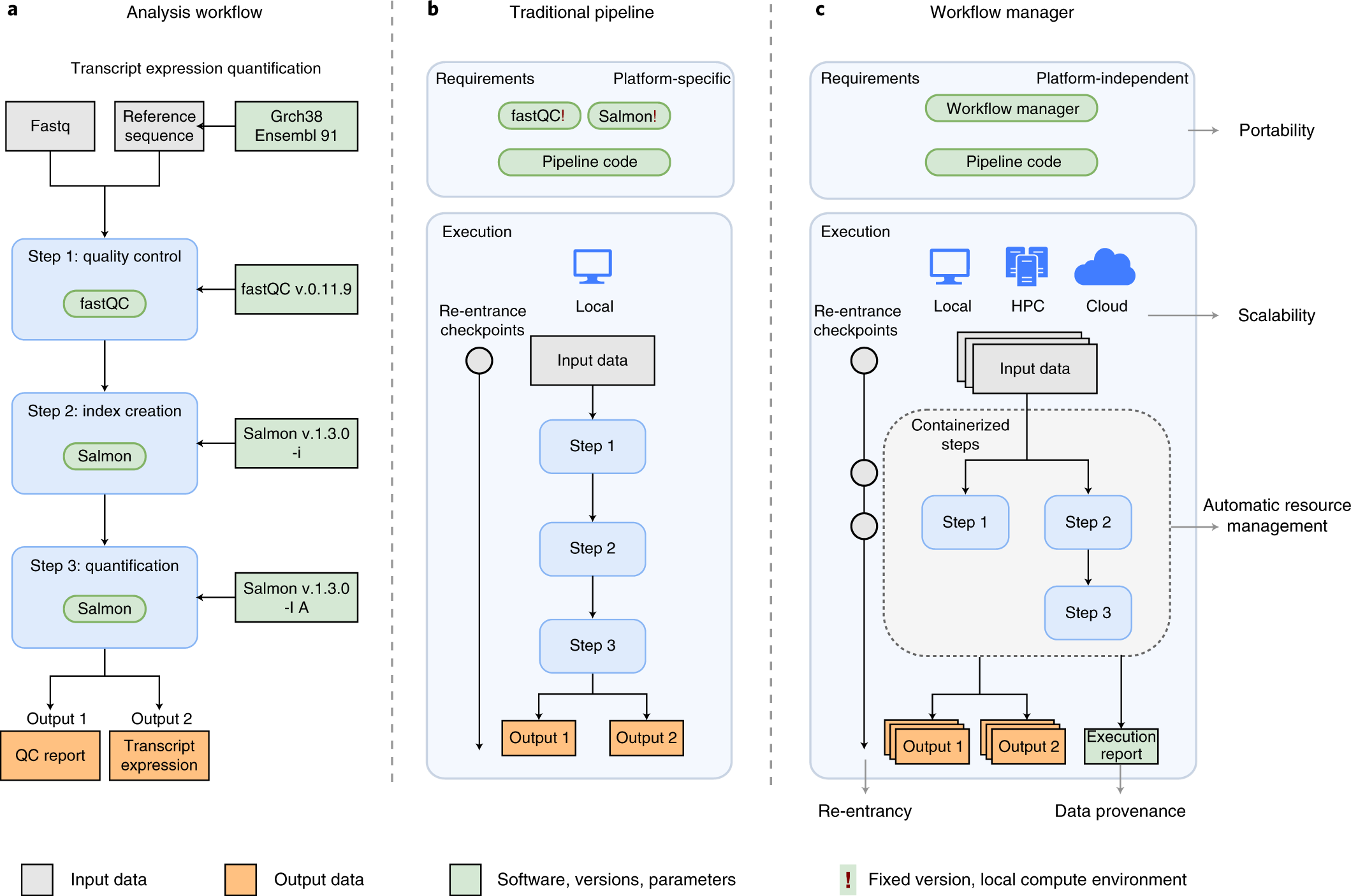
Modern bioinformatics research is powerful but deeply fragmented. A typical genomics study might require BLAST for sequence alignment, GATK for variant calling, DESeq2 for differential expression, and IQ-TREE for phylogenetic reconstruction — each tool installed separately, maintained independently, and connected through brittle shell scripts or manual file transfers. This fragmentation is not merely inconvenient; it is a reproducibility crisis. A 2023 survey found that fewer than 20% of published bioinformatics pipelines could be fully reproduced by independent groups, largely due to undocumented tool versions, missing dependencies, and environment-specific configurations.
The Model Context Protocol (MCP), originally developed as a standardised interface for connecting AI models to external tools and data sources, is now emerging as a transformative infrastructure layer for bioinformatics. By wrapping each tool as an MCP-compliant service with a unified API, MCP enables any client — whether an AI assistant, a workflow engine, or a researcher's browser — to invoke complex bioinformatics operations through a single, standardised interface.
Workflomics is the most comprehensive implementation of this vision to date: a central MCP server hub that connects 39+ open-source bioinformatics tools and 30 databases, enabling researchers to compose, execute, and benchmark multi-step analysis pipelines across 12 distinct biological scenarios.
What Is the Model Context Protocol?
The Model Context Protocol is an open standard that defines how software clients communicate with external tools and data sources. In the context of bioinformatics, MCP transforms each tool — BLAST, SAMtools, AlphaFold2, CRISPOR — from a standalone executable into a network-accessible service with a well-defined input/output schema. This means that instead of writing custom wrapper scripts for every tool combination, researchers can compose pipelines declaratively, describing what they want to compute rather than how each tool should be invoked.
The architectural advantages are significant. Docker containerisation ensures that every tool runs in an isolated, reproducible environment regardless of the host operating system. The unified API layer means that adding a new tool to the ecosystem requires only writing a single MCP wrapper, not re-engineering the entire pipeline. And because all tools communicate through the same protocol, benchmarking across tools becomes straightforward — you can compare STAR and Kallisto for RNA-seq quantification using identical inputs and standardised performance metrics without writing any additional code.
Workflomics: Architecture and Capabilities
Workflomics organises its 39+ integrated tools into 12 analysis scenarios, each representing a complete research workflow domain:
| Scenario | Key Tools | Primary Databases |
|---|---|---|
| DNA/RNA Analysis | BLAST, BWA, Trinity | GenBank, NCBI, Ensembl |
| Protein Analysis | MAFFT, HMMER | UniProt, PDB |
| Variant Calling | GATK, SAMtools, VEP | dbSNP, ClinVar, gnomAD |
| Metagenomics | Kraken2, MetaPhlAn | NCBI, Silva |
| Phylogenetics | IQ-TREE, BEAST2, MAFFT | TreeBASE, TimeTree |
| GWAS | PLINK, GCTA, REGENIE, SAIGE | GWAS Catalog, 1000 Genomes |
| Population Genetics | ADMIXTURE, PLINK | 1000 Genomes |
| Protein Structure | AlphaFold2, RoseTTAFold, ESMFold | AlphaFold DB, PDB |
| Protein Docking | AutoDock Vina, HADDOCK, Rosetta | BindingDB, ChEMBL |
| Gene Expression | DESeq2, edgeR, STAR, Kallisto | GEO, ArrayExpress, GTEx |
| Plasmid Design | Primer3, APE, pLannotate, SnapGene | Addgene |
| CRISPR Editing | CRISPOR, Cas-OFFinder, CRISPResso2 | Addgene |
The platform's visual workflow builder provides a drag-and-drop interface for composing multi-step pipelines, with pre-built templates for each of the 12 scenarios. Researchers can start from a template, customise parameters, and execute the pipeline with a single click — with real-time monitoring showing step-level logs, progress indicators, and result visualisations as the pipeline runs.
The Benchmark System: A Critical Innovation
Perhaps the most significant innovation in Workflomics is its integrated benchmark system. Bioinformatics tools are rarely evaluated on a level playing field. Published benchmarks often use different datasets, different parameter settings, and different hardware configurations, making direct comparison nearly impossible. Workflomics addresses this by running competing tools on identical inputs under controlled conditions and reporting standardised metrics across four dimensions: runtime, memory consumption, accuracy (measured against gold-standard reference datasets), and computational cost.
This benchmarking infrastructure has immediate practical value. A researcher choosing between STAR and Kallisto for RNA-seq quantification, or between AutoDock Vina and HADDOCK for protein docking, can consult Workflomics benchmarks to make an evidence-based decision rather than relying on anecdotal recommendations or outdated literature.
Implications for Biosecurity and Regulatory Science
From a biosecurity and regulatory perspective, the Workflomics MCP architecture offers capabilities that go beyond research efficiency. Regulatory agencies evaluating genetically modified organisms, novel biologics, or gene drive proposals require reproducible, auditable computational evidence. A pipeline executed through Workflomics generates a complete provenance record: every tool version, every parameter setting, every database query is logged and reproducible. This audit trail is precisely what regulatory frameworks such as the Cartagena Protocol on Biosafety and the WHO guidance on gene drive governance require.
Furthermore, the integration of CRISPR editing tools (CRISPOR, Cas-OFFinder, CRISPResso2) with population genetics tools (ADMIXTURE, PLINK) and phylogenetic reconstruction (IQ-TREE, BEAST2), creates a unified computational environment for the kind of ecological risk assessment that biosecurity experts need when evaluating gene drive proposals or horizontal gene transfer risks.
Getting Started
Workflomics is available at www.workflomics.com. The platform offers a free tier for researchers to explore the workflow builder and browse available pipelines. The MCP server hub can be connected to AI assistants and custom analysis environments, making it a foundational infrastructure layer for any research group working at the intersection of genomics, structural biology, and computational life sciences.
Reference
This post references the capabilities of Workflomics, a central MCP server hub for unified and benchmarked bioinformatics workflows. Workflomics connects 39+ open-source tools and 30 databases across 12 analysis scenarios. Visit www.workflomics.com to explore the platform.