The 2026 Annual Meeting of the American Association for Cancer Research revealed a scientific narrative that few observers had anticipated: the dramatic resurgence of spatial biology as the central platform for translational cancer research. After several years in which the field struggled with technical limitations, high costs, and questions about clinical utility, spatial biology has returned to the forefront of oncology — not through incremental improvement but through a fundamental convergence with artificial intelligence foundation models and multi-omics integration that is producing insights previously inaccessible to any single analytical approach.
Spatial biology refers to the family of technologies that measure molecular information — gene expression, protein abundance, metabolite concentrations, epigenetic modifications — while preserving the physical location of each measurement within a tissue sample. Unlike traditional bulk sequencing or even single-cell approaches, which dissociate tissue into individual cells and lose all spatial context, spatial methods reveal how cells are organised relative to one another, how they communicate through signalling networks, and how the architecture of tissue microenvironments influences disease progression and treatment response. In cancer research, this spatial context is not merely informative — it is often decisive, because the behaviour of a tumour cell depends critically on its neighbours, its distance from blood vessels, and its position within the broader immune landscape.
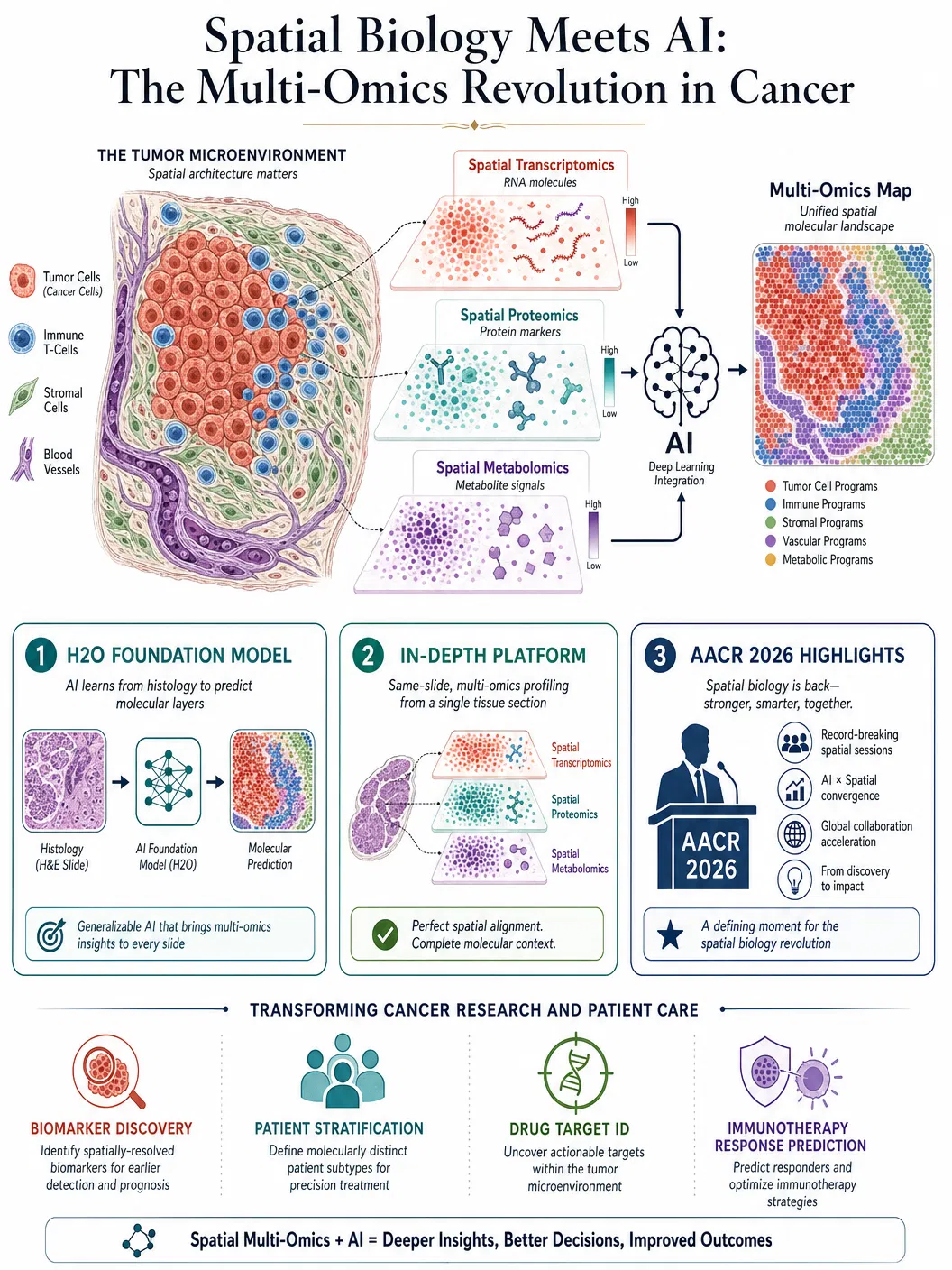
The key development showcased at AACR 2026 was the integration of spatial biology with AI foundation models — large neural networks pre-trained on massive datasets of tissue images and molecular profiles that can then be fine-tuned for specific analytical tasks. Bruker Spatial Biology presented multi-platform workflows demonstrating how high-fidelity spatial data from different measurement modalities — spatial transcriptomics, spatial proteomics, and spatial metabolomics — can be computationally integrated to produce comprehensive molecular maps of tumour microenvironments. These maps reveal not just which genes are expressed in which cells, but how the spatial organisation of gene expression patterns correlates with clinical outcomes such as treatment response, metastatic potential, and patient survival.
A particularly striking example was the H2O foundation model, described in a preprint published in April 2026, which bridges histopathology — the traditional microscopic examination of tissue that pathologists have performed for over a century — with spatial multi-omics data. The model is trained to predict spatial molecular profiles directly from standard histology images, effectively enabling researchers to extract multi-omics-level information from tissue slides that were prepared using conventional, inexpensive staining methods. If validated at scale, this approach could democratise access to spatial molecular insights by eliminating the need for expensive specialised assays, making precision oncology more accessible to hospitals and research institutions that lack cutting-edge spatial biology platforms.
The IN-DEPTH platform, published in a peer-reviewed article in April 2026, demonstrated another critical advance: same-slide spatial multi-omics integration. Rather than requiring separate tissue sections for different molecular measurements — a limitation that introduces technical variability and wastes precious clinical material — IN-DEPTH enables the measurement of multiple molecular layers from a single tissue section. This capability is particularly important for clinical applications, where biopsy material is often limited and every section counts. The platform revealed clinically relevant microenvironmental mechanisms in cancer that were invisible to any single-omics approach, underscoring the principle that biological complexity requires multi-dimensional measurement.
The convergence of spatial biology and AI is producing translational insights at a pace that has surprised even optimistic observers. As one leading researcher noted at AACR 2026, the combination of spatial data and AI foundation models is starting to make an "obvious difference" in translational research — a measured assessment from a community that is typically cautious about overstating progress. The difference is most apparent in three areas: biomarker discovery, where spatial AI models can identify complex tissue architecture patterns that predict treatment response more accurately than any single molecular marker; patient stratification, where spatial profiles enable the identification of patient subgroups that respond differently to immunotherapy based on the spatial organisation of their immune cells relative to tumour cells; and drug target identification, where spatial multi-omics reveals cell-cell interactions and signalling pathways that represent novel therapeutic vulnerabilities.
The implications for immunotherapy are particularly profound. The success or failure of checkpoint inhibitor therapy — the class of cancer drugs that has transformed oncology over the past decade — depends critically on the spatial relationship between immune cells and tumour cells within the microenvironment. Patients whose tumours are infiltrated by T cells in close proximity to cancer cells tend to respond well; patients whose immune cells are excluded from the tumour core, trapped at the periphery by physical or chemical barriers, tend to resist treatment. Spatial biology provides the tools to measure these relationships with unprecedented precision, and AI models can learn to predict treatment response from spatial patterns that are too complex for human pathologists to evaluate consistently.
The economic and infrastructural challenges of spatial biology remain significant. The technologies are expensive, the data volumes are enormous, and the computational requirements for AI-powered analysis are substantial. A single spatial transcriptomics experiment can generate terabytes of data, and the foundation models required to analyse that data demand GPU clusters that are beyond the reach of many academic laboratories. However, the trajectory of cost reduction in genomics — where the price of sequencing a human genome fell from three billion dollars to under two hundred dollars in two decades — suggests that spatial biology will follow a similar path, particularly as competition among platform providers intensifies and AI models become more efficient.
The broader significance of the spatial biology renaissance at AACR 2026 extends beyond cancer research. The same principles — preserving spatial context, integrating multiple molecular layers, and applying AI to extract patterns from high-dimensional data — are applicable to neuroscience, developmental biology, infectious disease, and virtually every field where tissue architecture matters. The convergence of spatial measurement and artificial intelligence is not merely a new tool for cancer researchers; it is a new paradigm for understanding how biological systems are organised in space and time. The insights emerging from this convergence are already reshaping clinical oncology, and their impact on the broader life sciences is only beginning to be felt.